

The necessity of internal reference gene screening
Endogenous control genes, also known as reference genes, are an important component of gene expression experiments. Using internal reference genes to normalize changes in target gene expression is currently the most accurate method to address potential biases caused by differences in sample processing and loading.
The reference gene must be stably expressed in all test samples, meaning that its expression level should not change between different tissue or cell sources, different treatments, different time points, or other experimental conditions.
Because the reference gene and the target gene are in the same sample preparation step, accurate quantification of the reference gene can homogenize the following differences that exist between different samples:
(i)Different quantities of starting materials(Variation in the amount of starting material)
(ii)Quality of starting materials(RNA/DNA quality)
(iii)Differences between RNA Extraction and cDNA Synthesis(RNA preparation &Reverse transcription efficiency)
Although any gene stably expressed under specified experimental conditions can be used as an internal reference gene, the most commonly selected types of internal references by researchers include constitutive housekeeping genes, such as cytoskeletal protein (ACTB) or ribosomal RNA (18S rRNA); Or genes involved in basic cellular functions, such as genes related to glycolysis pathways (GAPDH, PGK1) and protein folding (PPIA). At present, several common reference genes, including 18S rRNA, ACTB, and GAPDH, have been widely used to normalize differential expression, but they have not been experimentally validated.
However, a large number of studies have shown that these genes exhibit different expression levels in different tissues and under different experimental conditions, and should be used with caution [1]. Suzuki et al. [2] reported that in a certain year, over 90% of RNA transcription analyses in high impact journals used only one reference gene, with the most commonly used genes being 3-phosphoglyceraldehyde dehydrogenase (GAPDH), β - actin (ACTB), and 18S and 28S rRNA. However, some studies have indicated that there is a significant difference between β - actin and GAPDH, making them unsuitable as references for RNA transcription analysis [3]. Meanwhile, there are also reports that the abundance of 18S is too high for certain experiments, and using GAPDH may be superior to using 18S [4].
In the following figure, we tested the expression of 10 candidate reference genes in 6 different treatment samples and observed significant expression differences among most of the candidate reference genes in different treatment groups. Test data indicates that even when using "common" reference genes, their expression stability should still be considered. To avoid errors caused by changes in internal parameter expression due to experimental processing.
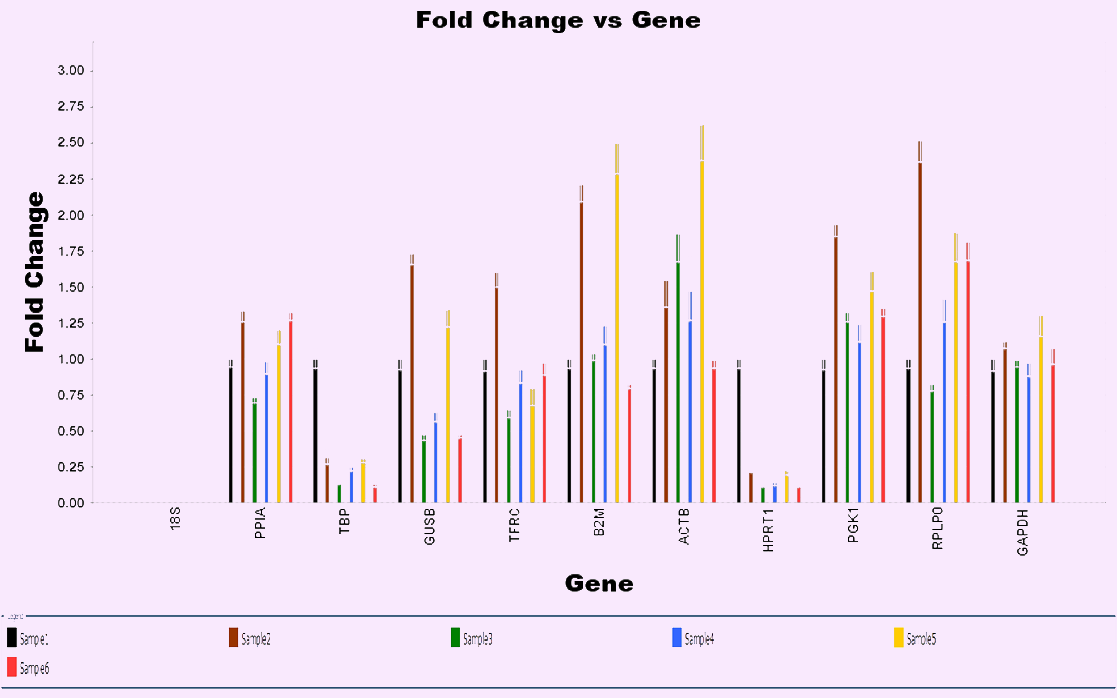
Figure 1. Expression of 10 candidate endogenous control genes in 6 samples with different treatments. Using 18S as control, the expression of 10 common candidate reference genes was studied, and the expression differences between different treatment samples were observed. It is suggested that even if we use “common” endogenous control, the expression stability should also be considered.
The best approach for this situation is to evaluate several candidate genes, as the ideal control for each experiment will depend on many variables, including the types of cells or tissues involved and the range of conditions to be tested. Certain housekeeping genes that encode proteins required for basic cellular functions are typically expressed at constitutive levels across a range of cell types and conditions, including disease states. Although these housekeeping genes can be good candidates for endogenous control and are worth considering, the expression of some classic housekeeping genes, such as β - actin and glyceraldehyde 3-phosphate dehydrogenase (GAPDH), varies greatly between tissue types. Before using housekeeping genes as endogenous controls in gene expression studies, it is necessary to detect the variability of housekeeping gene expression.
Internal Reference Screening Kit: The Key to Accurate Gene Expression Analysis
In the field of gene expression detection, the selection of reference genes is crucial. Inappropriate selection of reference genes can interfere with the accuracy of experimental results. To address this challenge, we have launched an internal reference screening kit.
Product Features:
Internal reference screening: Our reagent kit allows users to screen internal reference genes based on their research field, helping customers select the most suitable internal reference and improve the accuracy of experiments.
Wide selection: The internal reference Assay library contains dozens of validated genes, providing a wide range of choices to meet different research needs.
Optimize design: Each internal reference assay has been carefully designed and validated to ensure excellent performance under various experimental conditions.
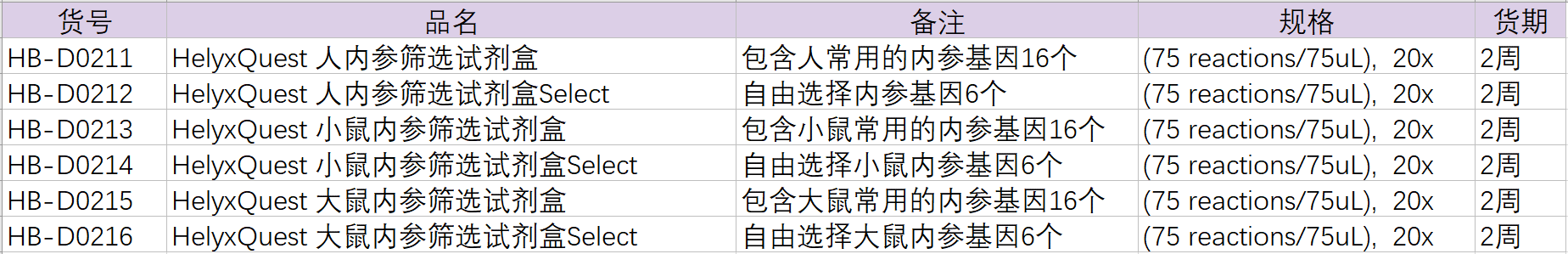
The following are the item numbers of the internal reference screening kit.
If you need more assistance or would like to learn more about product details, please feel free to contact our professional sales representative or customer service. We are always ready to provide you with personalized consultation and ordering services
References:
[1]Radonic A, Thulke S, Mackay IM et al. (2004) Guideline to reference gene selection for quantitative real-time PCR. Biochem Biophys Res Commun
[2]T Suzuki, P.J Higgins, D.R Crawford (2000)Control selection for RNA quantitation. Biotechniques
[3]S Selvey, E.W Thompson, K Matthaei, R.A Lea, M.G Irving(2001)
Beta-actin—an unsuitable internal control for RT-PCR. Mol. Cell Probes,
[4]K Gorzelniak, J Janke, S Engeli, A.M Sharma(2001),
Validation of endogenous controls for gene expression studies in human adipocytes and preadipocytes. Horm. Metab.
内参基因的选择列表如下:
Human | 18S rRNA | ACTB | B2M | GAPD | GUSB | HMBS | HPRT1 | IPO8 | PGK1 | POLR2A | PPIA | RPLPO | TBP | TFRC | UBC | YWHAZ |
Mouse | 18S rRNA | actb | b2m | gapdh | gusb | hmbs | hprt1 | ipo8 | pgk1 | polr2a | ppia | rplp2 | tbp | tfrc | ubc | ywhaz |
Rat | 18S rRNA | Actb | Arbp | B2m | Gapdh | Gusb | Hmbs | Hprt | Pgk1 | Ppia | Ppib | Rplp2 | Tbp | Tfrc | Ubc | Ywhaz |

Tel:400-998-6321
Postal code:215200
Address: 558 Fenhu Avenue, Lili Town, Wujiang District, Suzhou City, Jiangsu Province

